.: Core Research Activities in
Reconfigurable Computing Laboratory
|
5.
Many-Core Architecture Exploration for Scientific Applications (with
five active projects)
A)
iPlant Genotype-to-Phenotype (iPG2P) Grand Challenge Project ( Gregory
Streimer)
Funded
by: The iPlant Collaborative. The iPlant Collaborative is funded
by a grant from the National Science Foundation Plant Cyberinfrastructure
Program (#EF-0735191).
Elucidating
the relationship between plant genotypes and the resultant phenotypes
in complex (e.g., non-constant) environments is one of the foremost
challenges in plant biology. Plant phenotypes are determined by
often intricate interactions between genetic controls and environmental
contingencies. In a world where the environment is undergoing rapid,
anthropogenic change, predicting altered plant responses is central
to studies of plant adaptation, ecological genomics, crop improvement
activities (ranging from international agriculture to biofuels),
physiology (photosynthesis, stress, etc.), plant development, and
many many more. 
RCL
is participating in the NSF Funded
iPlant Collaborative by developing computationally intensive
algorithms to run on NVIDIA GPUs for relating genotype to phenotype
in complex environments.
High
throughput data analysis workflow in life sciences are typically
composed of iterative tasks that can potentially leverage the architectural
benefits afforded by GPGPUs to improve overall performance. The
compute bottlenecks in the workflow are interspersed and very often
are the rate limiting factors, it is important to consider the overall
execution environment and how GPGPU-based approaches can integrate
with the existing workflow. In collaboration with Prof. Steve Welch
(iPG2P) and designated working groups (Statistical Inference, Next
Gen), suitable candidate applications (algorithms) for exploration
will be identified. These identified application will involve solving
N equations with N unknowns. Matrix reduction and coefficient calculations
with large size vectors involve recurring sequences of instructions
(multiply-subtract, multiply-add, etc) within the loop level executions.
Traditional clusters can run these loops concurrently;
however, for largeN we expect GPGPUs to be a bettermatch to the
large number of fine-grain computations. The team at the University
of Arizona (Nirav Merchant, Arizona Research Lab, Prof. Ali Akoglu,
ECE), and (Prof. David Lowenthal, Computer Science) has been formed
to study the following aspects of GPGPUs:
- Evaluate
the effectiveness of Open CL and CUDA in developing GPGPU programs.
- Develop
architectural and application-level models to determine which
portions of a program to offload to the GPGPU.
- Explore
ways to restructure the program architecture in order to exploit
the unique memory hierarchy of the GPU and its processing power
with hundreds of processing cores.
- Demonstrate
the scalability of the developed algorithms to multiple GPUs.
- Quantify
the speed-up achieved with CUDA environment by performance comparison
against CPU based parallel system.
Publications:
- Peter
Bailey, Tapasya Patki, Gregory M. Striemer, Ali Akoglu, David
Lowenthal, Peter Bradbury, Matthew Vaughn, Liya Wang, and Stephen
Goff, "Quantitative Trait Locus Analysis Using a Partitioned
Linear Model on a GPU Cluster," IEEE International Workshop
on High Performance Computational Biology, HiCOMB'12, May 2012,
Shanghai, China.
- Stephen
A Goff, Matthew Vaughn, Sheldon McKay, Eric Lyons, Ann Stapleton,
Damian Gessler, Naim Matasci, Liya Wang, Matthew Hanlon, Andrew
Lenards, Andy Muir, Nirav Merchant, Sonya Lowry, Stephen Mock,
Matthew Helmke, Adam Kubach, Martha Narro, Nicole Hopkns, David
Micklos, Uwe Hilgert, Michael Gonzales, Chris Jordan, Edwin Skidmore,
Rion Dooley, John Cazes, Robert McLay, Zhenyuan Lu, Shiran Pasternak,
Lars Koesterke, William H. Piel, Ruth Grene, Christos Noutsos,
Karla Gendler, Xin Feng, Chunlao Tang, Monica Lent, Seung-Jin
Kim, Kristian Kvilekval, B. S. Manjunath, Val Tannen, Alexandros
Stamatakis, Michael Sanderson, Stephen W. Welch, Karen A. Cranston,
Pamela Soltis, James Leebens-Mack, Michael J. Donoghue, Edgar
P. Spalding, Todd J. Vision, Christopher R. Myers, David Lowenthal,
Brian J. Enquist, Brad Boyle, Ali Akoglu, Greg Andrews, Sudha
Ram, Doreen Ware, Lincoln Stein, and David Stanzione, "The
iPlant collaborative: cyberinfrastructure for plant biology,"
Frontiers in Plant Science, vol. 2, no.34, pp. 1-16, 2011.
- Gregory
M. Striemer, Ali Akoglu, David Lowenthal, Peter Bradbury, Liya
Wang, Matthew Vaughn, Stephen Goff, "Relating Genotypes to
Phenotypes in Complex Environments: Generalized Linear Model (GLM)
Based Quantitative Trait Locus (QTL) Analysis", NVIDIA GPU
Technology Conference, October 20-23, 2010, San Jose, CA
B)
Parallel Implementation of Irregular Terrain Model on NVIDIA Graphic
Processing Units and IBM Cell Broadband Engine
Funded
by: United States Army Battle Command Battle Laboratory - Huachuca
(BCBL-H)
Students:
Yang Song
Advances
in digital communication technologies have enabled sophisticated
devices that are adaptive to changes in the surrounding environment
for broadband connectivity. In telecommunications, white spaces
refer to vacant frequency bands between licensed broadcast channels
or services like wireless microphones. After the transition to digital
TV in the U.S. in June 2009, the amount of white space exceeded
the amount of occupied spectrum even in major cities. Utilization
of white spaces for digital communications requires propagation
loss models to detect occupied frequencies in near real-time for
operation without causing harmful interference to a DTV signal,
or other wireless systems operating on a previously vacant channel.
However, signal propagation is an enigmatic phenomenon whose properties
are usually difficult to predict, especially at Very High Frequency
(VHF), Ultra High Frequency (UHF), and Super High Frequency (SHF).
Complications are further amplified by the clutter of hills, trees,
houses, and the ever-changing atmosphere provide scattering obstacles
with sizes of the same order of magnitude as the wavelength. 
In
order to address these difficulties, a variety of propagation models,
either theoretically or experimentally based, have been proposed
to describe how the physical world affects the flow of electromagnetic
energy. For example, the Irregular Terrain Model (ITM), also known
as the Longley-Rice model, is used to make predictions of radio
field strength based on the elevation profile of terrains between
the transmitter and the receiver. In practice, the terrain features
that characterize the physical environment are tremendously diverse.
This contributes to the amount of data required to represent the
terrain. Additionally, radio transmitters have wide variations in
range capabilities, which results in a more dynamical frequency
coverage. Hence, ITM needs to involve many iterations of environmental
characterizations. This intensive repeated serial workload poses
as the main barrier for common mainstream microprocessors in responding
to the near real-time demand for evaluating TV service coverage,
and interference based on the real and hypothetical television transmitters
in accordance with the Federal Communications Commission (FCC) rules.
ITM
predicts long-term median transmission loss of a radio signal based
on atmospheric and geographic conditions. Due to constant changes
in terrain conditions and variations in radio propagation, there
is a pressing need for computational resources to run hundreds of
thousands of transmission loss calculations per second. Multicore
processors like the NVIDIA Graphics Processing Unit (GPU) and IBM
Cell Broadband Engine (BE) offer improved performance over mainstream
microprocessors for ITM. In this project, we compare architectural
features of the Tesla C870 GPU and Cell BE, and evaluate the effectiveness
of various architecture-specific optimizations and parallelization
strategies while mapping ITM onto these platforms. We evaluate the
GPU implementations which utilize both global and shared memories
along with fine-grained parallelism. We then compare the GPU performance
with Cell BE implementations which utilize direct memory access,
double buffering and partial SIMDization.
Publications:
- Yang
Song and Ali Akoglu, "Parallel implementation of the irregular
terrain model (ITM) for radio transmission loss prediction using
GPU and Cell BE processors," IEEE Transactions on Parallel
and Distributed Systems, (TPDS), vol. 22, no. 8, pp. 1276-1283,
2011.
- Yang
Song, Ali Akoglu, "Parallel Implementation of Irregular Terrain
Model on IBM Cell Broadband Engine", 23rd IEEE International
Parallel and Distributed Processing Symposium, May 25-29, 2009,
Rome, Italy
C)
Sequence Alignment (Student: Gregory Streimer and Yang song)
A common
problem posed in computational biology is the comparison of protein
sequences with unknown functionality to that of a set of known protein
 sequences to detect functional similarities. The most sensitive
algorithm for searching similarities is the Smith-Waterman algorithm,
however it is also the most time consuming. With an ever increasing
size in protein and DNA databases, searching them becomes more difficult
with exact algorithms such as Smith-Waterman. This research explores
reducing the computational time of the algorithm by utilizing multi-core
technologies. Currently we are examining the potential of Compute
Unified Device Architecture (CUDA) using the Nvidia C870 graphics
processing unit (GPU). This card has massively parallel computational
capabilities, which makes it ideal for high-performance scientific
computing. The C870 has sixteen multi-processors, each containing
eight streaming processors which individually operate at 1.35 GHz.
Each multi-processor is capable of launching up to 768 threads at
once, and each thread can run a Smith-Waterman alignment on a different
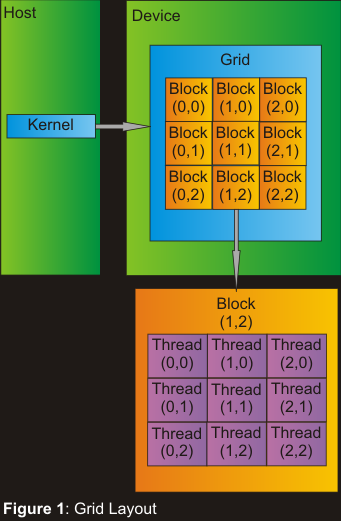
sequence pair. On the GPU, the threads are organized in blocks within
a grid of blocks which are launched from the kernel. This can be
seen in Figure 1.
sequences to detect functional similarities. The most sensitive
algorithm for searching similarities is the Smith-Waterman algorithm,
however it is also the most time consuming. With an ever increasing
size in protein and DNA databases, searching them becomes more difficult
with exact algorithms such as Smith-Waterman. This research explores
reducing the computational time of the algorithm by utilizing multi-core
technologies. Currently we are examining the potential of Compute
Unified Device Architecture (CUDA) using the Nvidia C870 graphics
processing unit (GPU). This card has massively parallel computational
capabilities, which makes it ideal for high-performance scientific
computing. The C870 has sixteen multi-processors, each containing
eight streaming processors which individually operate at 1.35 GHz.
Each multi-processor is capable of launching up to 768 threads at
once, and each thread can run a Smith-Waterman alignment on a different
sequence pair. On the GPU, the threads are organized in blocks within
a grid of blocks which are launched from the kernel. This can be
seen in Figure 1.
Current
limitations on the GPUs memory can constrain how much data can be
processed at any given time, so exploiting different memory resources
to their fullest potential for Smith-Waterman is also a part of
this research. Due to some of the memory restrictions, our implementation
utilizes the system's cached constant memory for storage of the
substitution matrices, as well as the user's query sequence. This
allows for faster access to these highly utilized values through
the use of an extremely efficient cost function. A major advantage
of our implementation over other GPU implementations is the fact
that we do not place restrictions on the length of database sequences,
so a database can truly be searched in its entirety. This work will
be benchmarked against another known GPU implementation, as well
as commonly used serial and multi-threaded CPU implementations.
Figure 2 displays our basic program's flow
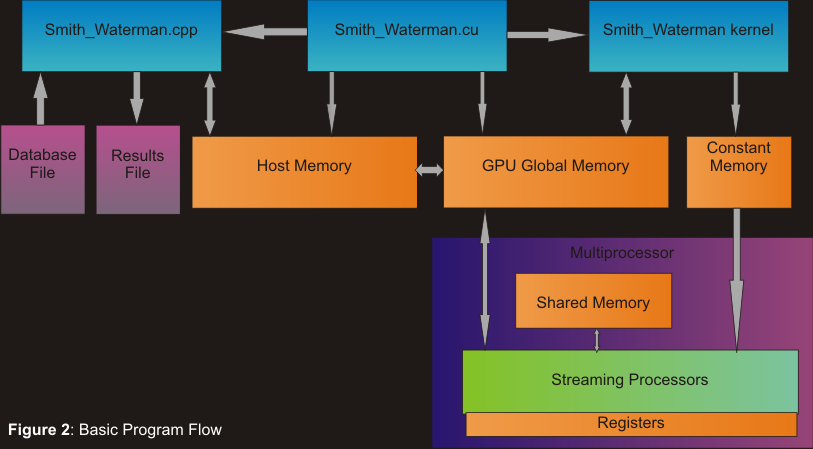
Publications:
- Khaled
Benkrid, Ali Akoglu, Cheng Ling, Yang Son, Ying Liu, and Xiang
Tian, "High performance biological pairwise sequence alignment:
FPGA vs. GPU vs. Cell BE vs. GPP," International Journal
of Reconfigurable Computing, special issue on "High Performance
Reconfigurable Computing," accepted for publication, 2011.
- Yang
Song,
Gregory M. Striemer and Ali Akoglu, "Performance Analysis
of IBM Cell Broadband Engine on Sequence Alignment", IEEE
NASA/ESA Conference on Adaptive Hardware and Systems (AHS 2009),
July 2009, San Francisco
- Greg
Streimer, Ali Akoglu, "Sequence Alignment with GPU: Performance
and Design Challenges", 23rd IEEE International Parallel
and Distributed Processing Symposium, May 25-29, 2009, Rome, Italy
This
study is posted on CUDA
Zone
Source
Code
fror Smith-Waterman Algorithm for use on an NVIDIA GPU using CUDA
(posted 6/16/2009)
D)
Cardiac Simulation on Multi-GPU Platform (Student Venkata Krishna
Nimmagadda)
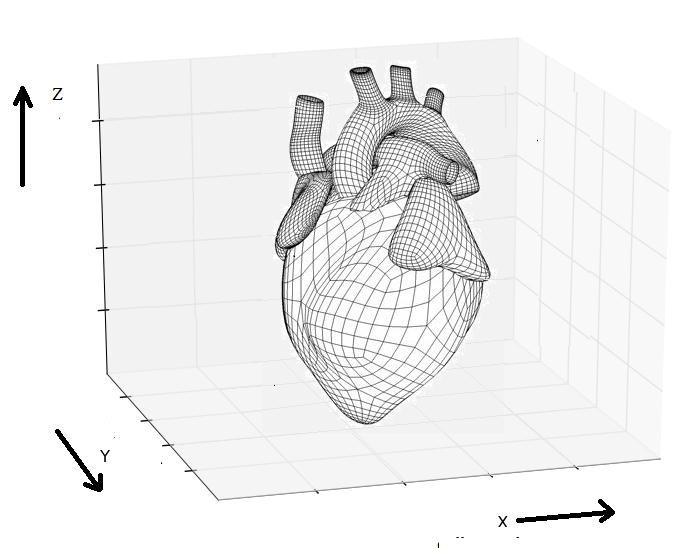  Cardiac
Bidomain Model is a popular approach to study electrical behavior
of tissues and simulate interactions between the cells by solving
partial differential equations. The iterative and data parallel
model is an ideal match for the parallel architecture of Graphic
Processing Units (GPUs). In this study, we evaluate the effectiveness
of architecture-specific optimizations and fine grained parallelization
strategies, completely port the model to GPU, and evaluate the performance
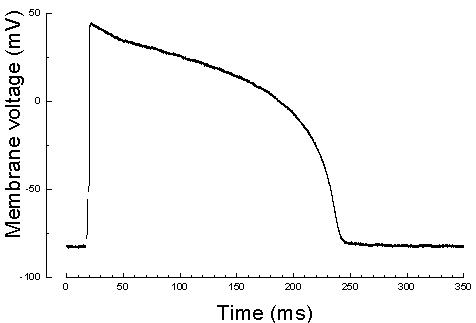
of single-GPU and multi-GPU implementations. Simulating one action
potential duration (350 msec real time) for a 256x256x256 tissue
takes 453 hours on a high end general purpose processor, while it
takes 664 seconds on a four-GPU based system including the communication
and data transfer overhead. This drastic improvement (a factor of
2460x) will allow clinicians to extend the time-scale of simulations
from milliseconds to seconds and minutes; and evaluate hypotheses
in shorter amount of time that was not feasible previously. Cardiac
Bidomain Model is a popular approach to study electrical behavior
of tissues and simulate interactions between the cells by solving
partial differential equations. The iterative and data parallel
model is an ideal match for the parallel architecture of Graphic
Processing Units (GPUs). In this study, we evaluate the effectiveness
of architecture-specific optimizations and fine grained parallelization
strategies, completely port the model to GPU, and evaluate the performance
of single-GPU and multi-GPU implementations. Simulating one action
potential duration (350 msec real time) for a 256x256x256 tissue
takes 453 hours on a high end general purpose processor, while it
takes 664 seconds on a four-GPU based system including the communication
and data transfer overhead. This drastic improvement (a factor of
2460x) will allow clinicians to extend the time-scale of simulations
from milliseconds to seconds and minutes; and evaluate hypotheses
in shorter amount of time that was not feasible previously.
Publications:
- Venkata
Krishna Nimmagadda, Ali Akoglu, Salim Hariri, and Talal Moukabary,
"Cardiac simulation on multi-GPU platform," Journal
of Supercomputing, vol 59, no. 3, pp. 1360-1378, 2011. http://dx.doi.org/10.1007/s11227-010-0540-x
E)
Numerical solution of partial differential equations (PDEs) with
GPU
Funded
by: Air Force, Contract # FA9550-10-C-0104
Numerical
solution of partial differential equations (PDEs) is one of the
essential components in design, manufacturing and analysis of machines
and structures. There are different approaches for solving PDEs.
For example, PDEs governing structural problems are often solved
using the finite element method (FEM) whereas finite difference
method (FDM) and finite volume method (FVM) are commonly utilized
for solution of PDEs governing fluid dynamics. For more than half
a century, aeronautics industry has been heavily relying on such
analyses for the advancement of science. Our goal is to design and
develop a framework and a library of algorithms optimized for the
GPU architecture and targeted for PDE based problems. These algorithms
will address problems relevant and important to aeronautics industry,
which include solid mechanics, heat transfer and fluid dynamics.
Publications:
- Yoon
Kah Leow, Ali Akoglu, Ibrahim Guven and Erdogan Madenci, "High
performance linear equation solver using NVIDIA graphical processing
units," IEEE NASA/ESA Conference on Adaptive Hardware and
Systems (AHS), San Diego, CA, Jun. 6-9, 2011, pp. 367-374.
|




